In February 2014, I did a series of posts on my deep maternal ancestors, identified through a test of mutations on my mitochondrial DNA (mtDNA) which is inherited only from the mother. This test was carried out by Ancestry.com, who have since discontinued tests of mtDNA and Y-chromosome DNA. Costs of DNA tests have dropped dramatically since then, and late last year I ordered an mtDNA test from FamilyTreeDNA (www.familytreedna.com) which carried out a full sequencing of the mitochondrial DNA.
As well as the DNA that makes up the chromosomes in the nuclei of our cells, we also have another type of DNA called mitochondrial DNA (mtDNA). The mitochondria are organs located outside the cell nucleus which convert sugars into energy. Mitochondria have a small circular loop of DNA, containing only approximately 16,569 base pairs in humans. The circular mtDNA is similar to the DNA of bacteria, and it is thought that mitochondia evolved from symbiotic bacteria that were once free living.
In humans, as in other higher organisms, a DNA molecule consists of two strands that wrap around each other in the well known double helix structure. Each strand contains a linear arrangement of four different bases: adenine (A), thymine (T), guanine (G), and cytosine (C). Specific sequences of three bases make a DNA “word” which codes for an amino acid. A string of DNA “words” strung together in a sequence provides the instructions to make a particular protein. The base-pairs on the circular mtDNA loop are numbered from 1 to 16,569 and different portions of the loop have been given different names as shown in the following diagram. The first and second portions, called hypervariable control region 1 (HRV1) and hypervariable control region 2 (HRV2) are regions of the mtDNA that accumulate mutations (changes in base-pairs) relatively quickly and thus tend to be hyper-variable between people who are not closely related. The third portion, the coding region, accumulates far fewer changes and contains the base-pair sequences for mitochondrial genes [1].

mtDNA molecule (Credit: Debbie Parker Wayne)
One type of mutation, called a single nucleotide polymorphism (SNP), is a single base pair in a DNA sequence that has been replaced by a different base pair. My mtDNA test results identified 68 SNPs, 13 in HRV1, 17 in HRV2, and 38 in the coding region through comparison of my base sequences with those of the Reconstructed Sapiens Reference Sequence (RSRS). The RSRS is a recent effort to reconstruct a single ancestral genome for all living humans, using both a global sampling of modern human samples and samples from ancient hominids. It was introduced in early 2012 as a replacement for the rCRS (revised Cambridge Reference Sequence).
Mitochondrial DNA is inherited from the mother alone, rather than being inherited from the father and the mother. Additionally, recombination (or crossing over) does not occur in mitochondrial DNA (mtDNA). For both of these reasons, the sequence of mitochondrial DNA stays largely the same over generations, and thus is a useful tool for looking at maternal ancestry.
Over a period of nearly 180,000 years, SNPs have steadily accumulated on different human mtDNA molecules being passed down from mothers to their children. SNPs thus provide a cumulative dossier of our own maternal prehistory. We can use these mutations to reconstruct a genetic tree of mtDNA, because each new SNP in a prospective mother’s ovum will be transferred in perpetuity to all her descendants down the female line.
When a group of people share similar SNPs, they are part of the same “haplogroup”. For example, over 95% of native-born Europeans fit into seven haplogroups, which in turn derive from an older haplogroup that arose in the Middle East. Other regions of the world are associated with different haplogroups. Each of these groups trace their maternal ancestry back to just one woman, the common maternal ancestor of everyone in her haplogroup, or clan. Not everyone in the same clan has exactly the same mtDNA, because DNA accumulates additional mutations gradually over the generations. However, everyone in the clan shares a set of common mutations, which are the signature of the mtDNA of their founding maternal ancestor.
By averaging the numbers of mutations found in the mtDNA of modern members of a haplogroup, and knowing the average mutation rate for mtDNA, as well as the dating of ancient human remains whose mtDNA has been sequenced, it is possible to estimate how old each clan is, or in other words, when the common maternal ancestor of the clan lived (see below). By studying features of the geographical distribution of their present-day descendants, as well as the locations of ancient human remains whose mtDNA has been sequenced, it is possible to work out where they most likely lived as well. Generally speaking, the likely geographic origin for a clan is not necessarily the place where it is most common today, but the place where it is the most varied.
It is thus possible to trace migration routes by observing the branching points in an ancestral map containing all known haplogroups (see map below). Mitochondrial DNA in humans of African origin show the most diversity in the world. This supports the concept that ancient humans first existed in Africa and stayed in Africa for thousands of years. The humans who left Africa around 70,000 years age took two major routes, to Asia (haplogroup M) and to Europe (haplogroup N).
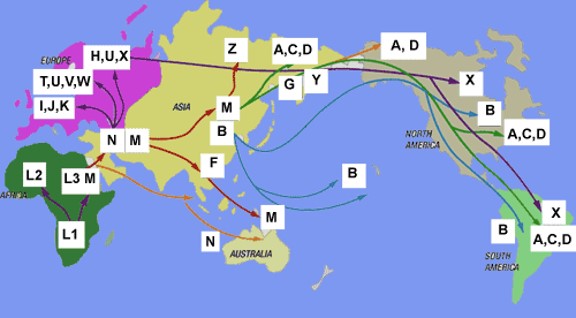
Migration routes of human beings dating back to 170,000 years ago. All humans originated in Africa and migrated out, branching into the two main out of Africa haplogroups, M and N. Individuals in haplogroup M headed west to Asia and later to the Americas, while haplogroup N moved into Europe.
The clan mothers were not the only people alive at the time, of course, but they were the only ones to have direct maternal descendants living right through to the present day. The other women around, or their descendants, either had no children at all or had only sons, who could not pass on their mtDNA. And, of course, the clan mothers had ancestors themselves. Everyone alive on the planet today can trace their maternal ancestry back to just one woman, the founder of haplogroup L. According to recent studies, she lived in Africa nearly 180,000 years ago and is known as “Mitochondrial Eve” (see my previous post mitochondrial eve )
Age Estimates
The age estimates for haplogroup founders in my 2014 posts were based on estimating average mutation rates for mtDNA SNPs, assuming an average constant mutation rate [2, 3]. There have been several more recent studies that have updated these estimates using not only samples from living humans but also analyses of mtDNA from ancient humans.
My previous post on mitochondrial Eve used dates from Behar et al. 2012 [4]. Comparing these with the dates given on the FamilyTreeDNA site and on the site snpTracker, as well as as two other more recent papers [5, 6], there is considerable variation in some of these. I revised the date for mitochondrial Eve based on the average of the dates from Behar et al [4] and Fu et al [5], and the dates for L3 and U based on the average of the dates from Behar et al [4], Fu et al [5] and Soares et al [6]. Dates from these three papers were reasonably consistent. Dates for other haplogroups from Behar et al were adjusted accordingly. 90% uncertainty ranges were estimated using the relative uncertainty ranges Behar et al. [4]. Because of the random nature of individual mutations (they may not occur at exactly the average time to next mutation), there is an uncertainty range around dates (which is also statistically estimated).
The following map summarizes the revised timeline for the migration of the ancestors of maternal haplogroup U5 out of Africa.

Ancestral migration path of maternal ancestors for haplogroup U5
In following posts, I will update the previous posts summarizing my maternal haplogroup ancestors and placing them in the context of the human expansion out of Africa and across Europe, as well as the context of the ice ages and the evolution of human cultures.
References
[1] Blaine T. Bettinger (2016). The Family Tree Guide to DNA Testing and Genetic Genealogy. Family Tree Books: Cincinnati.
[2] Behar D, Villems R, Soodyall H, Blue-Smith J, Luisa Pereira L, Ene Metspalu E, Rosaria Scozzari R, Heeran Makkan H, Shay Tzur, David Comas, Jaume Bertranpetit, Lluis Quintana-Murci, Chris Tyler-Smith, R. Spencer Wells, Saharon Rosset (2008). The Dawn of Human Matrilineal Diversity. American Journal of Human Genetics, 82(5):1130-1140.
[3] Soares P, Ermini L, Thomson N, Mormina M, Rito T, Röhl A, Salas A, Oppenheimer S, Macaulay V, Richards MB (2009). Correcting for purifying selection: an improved human mitochondrial molecular clock American Journal of Human Genetics, 84(6):740-59.
[4] Behar D, van Oven M, Rosset S, et al. A “Copernican” Reassessment of the Human Mitochondrial DNA Tree from Its Root. Am J Hum Genet. 2012;90(5):936. doi:10.1016/j.ajhg.2012.04.007 Open ArchiveDOI:https://doi.org/10.1016/j.ajhg.2012.03.002
[5] Fu Q, Mittnik A, Johnson PLF, et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr Biol. 2013;23(7):553–559. doi:10.1016/j.cub.2013.02.044 https://www.cell.com/current-biology/fulltext/S0960-9822(13)00215-7?code=cell-site
[6] Soares S, Alshamali F, Pereira JB, Fernandes V, Silva NM, Afonso C, Costa MD, Musilová E, Macaulay V, Richards MB, Černý V, Pereira L, The Expansion of mtDNA Haplogroup L3 within and out of Africa, Molecular Biology and Evolution, Volume 29, Issue 3, March 2012, Pages 915–927, https://doi.org/10.1093/molbev/msr245

Pingback: Our Asgardian ancestors | Mountains and rivers
Pingback: My maternal ancestors – from Eve via ice age Europe to Victorian England | Mountains and rivers
Pingback: A Tale of 2 Kidneys and Mitochondrial DNA – 'This Side of Life'